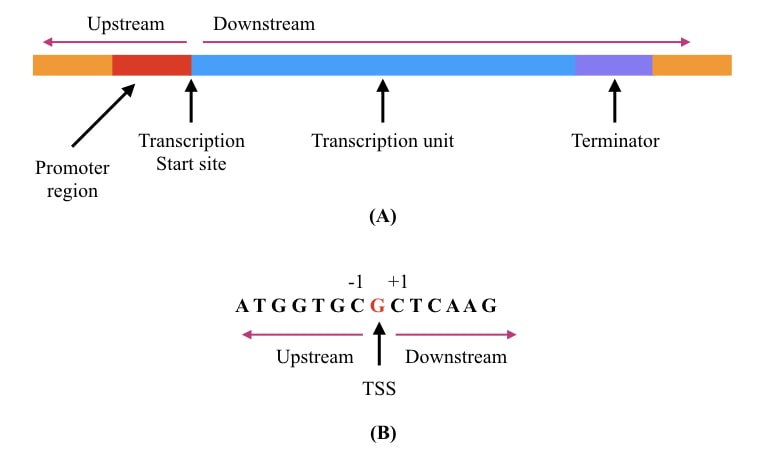
Global mapping transcriptional start sites revealed both transcriptional and post-transcriptional regulation of cold adaptation in the methanogenic archaeon Methanolobus psychrophilus | Scientific Reports

Histone modification enrichment in TSS (transcription start site) and gene body regions and its relationship with gene expression in 4 categories.

The Promoter Region of the MDR1 Gene Is Largely Invariant, but Different Single Nucleotide Polymorphism Haplotypes Affect MDR1 Promoter Activity Differently in Different Cell Lines | Molecular Pharmacology

Regulating retrotransposon activity through the use of alternative transcription start sites | EMBO reports
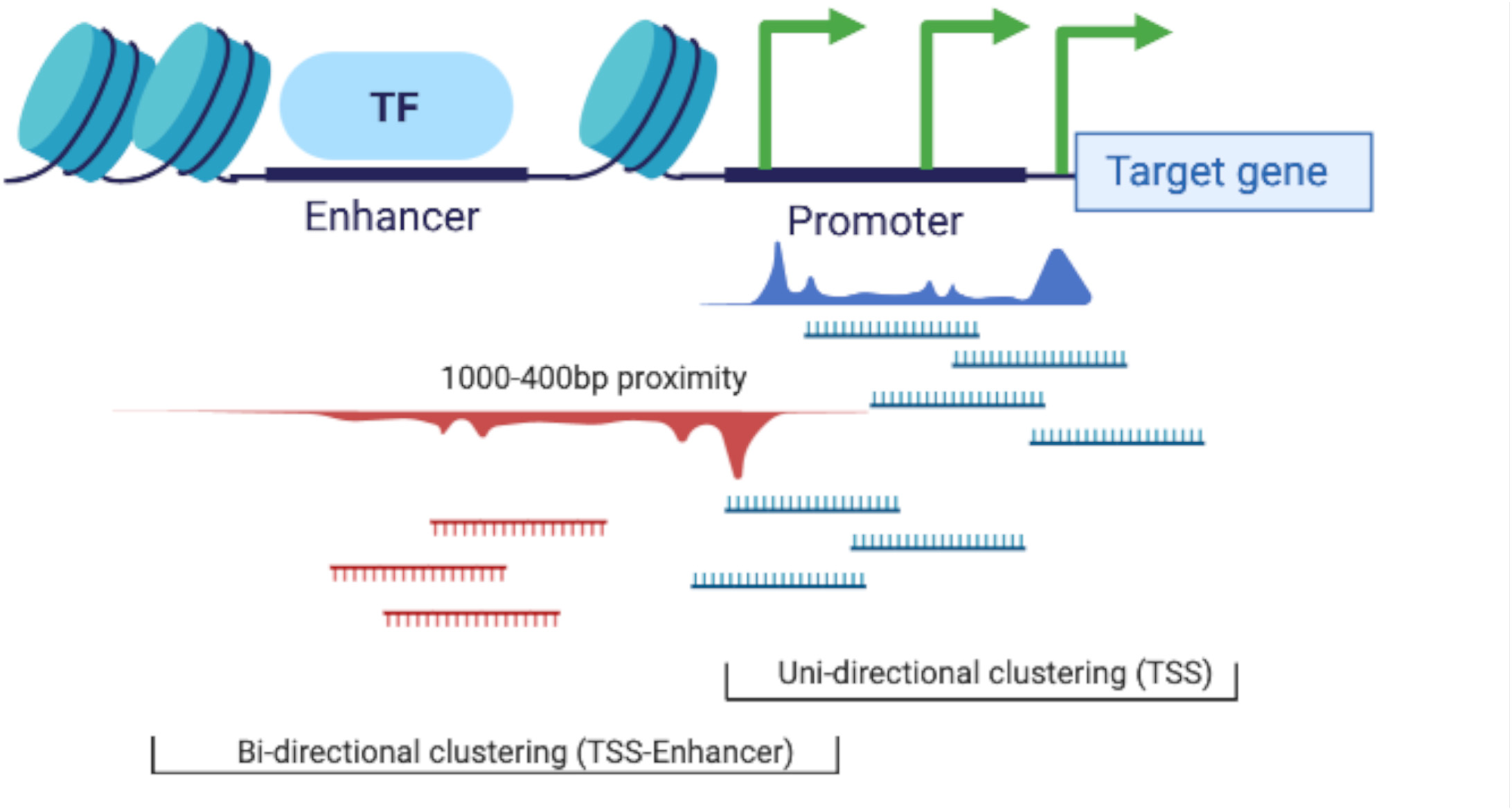
Frontiers | Global Analysis of Transcription Start Sites in the New Ovine Reference Genome (Oar rambouillet v1.0)

Biomolecules | Free Full-Text | AP-TSS: A New Method for the Analysis of RNA Expression from Particular and Challenging Transcription Start Sites

Where To Begin? Mapping Transcription Start Sites Genome-Wide in Escherichia coli | Journal of Bacteriology
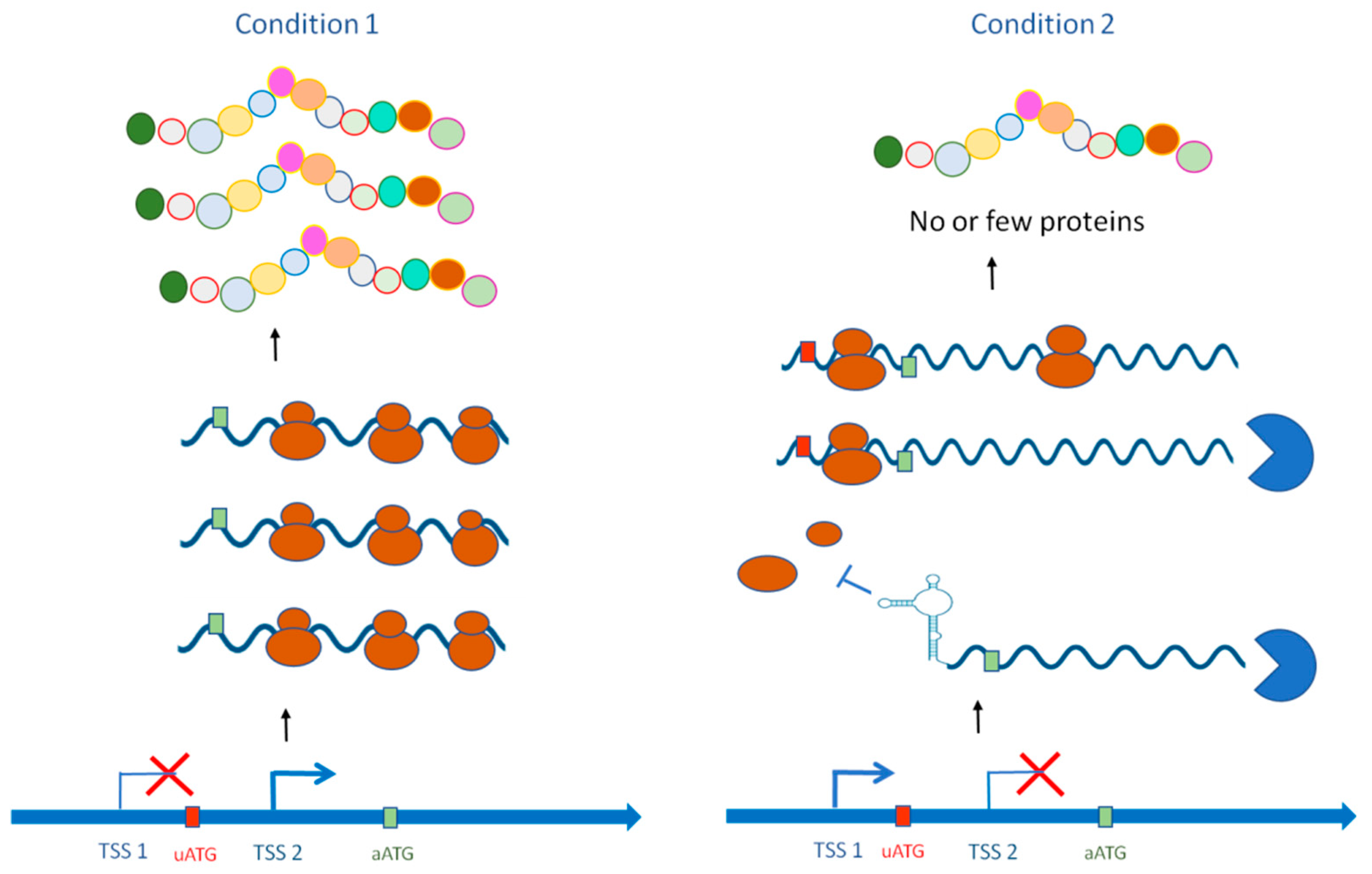
JoF | Free Full-Text | Alternative Transcription Start Site Usage and Functional Implications in Pathogenic Fungi
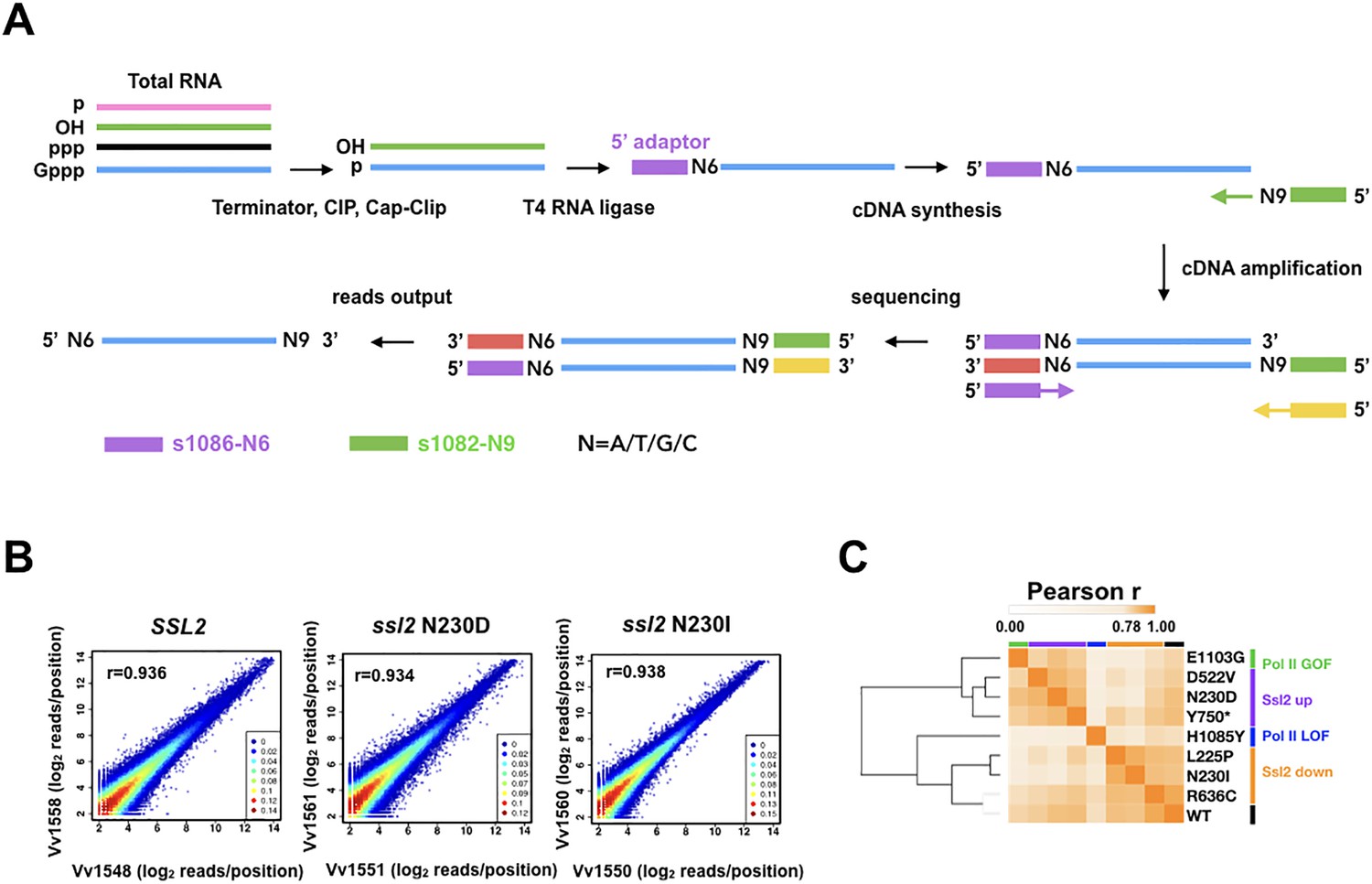
Figures and data in Ssl2/TFIIH function in transcription start site scanning by RNA polymerase II in Saccharomyces cerevisiae | eLife
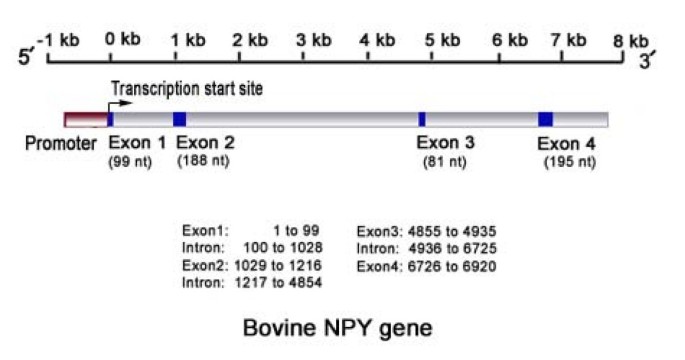

Mapping of the transcription start site (TSS) and identification of SNPs in the bovine neuropeptide Y (NPY) gene | BMC Genomic Data | Full Text

ARF-TSS: an alternative method for identification of transcription start site in bacteria | BioTechniques
Characterization of the Regulatory Region of the Zebrafish Prep1.1 Gene: Analogies to the Promoter of the Human PREP1 | PLOS ONE